Study Protocol / Open Access
DOI: 10.31488/bjcr.212
Comparison of Oral Microbiome in Patients with and Without Oral Squamous Cell Carcinoma: Study Protocol for a Prospective Case- Control Trial
Hagens ERC*1,2, Kiewiet CC1,2, Kessler PAWH1,2, Coppen C3, Savelkoul PHM4, Vaassen LAA1,2
1.Department of Cranio-Maxillofacial Surgery, Head and Neck Surgery, Maastricht University Medical Center, 6202 AZ Maastricht, The Netherlands
2.GROW School for Oncology and Developmental Biology, Maastricht University Medical Center, 6202 AZ Maastricht, The Netherlands
3.Department of Oral and Maxillofacial Surgery, Radboudumc, Nijmegen, The Netherlands
4.Department of Medical Microbiology, Infectious Diseases & Infection Prevention, School of Nutrition and Translational Research in Metabolism (NUTRIM), Maastricht University Medical Center, 6202 AZ Maastricht, The Netherlands
*Corresponding author:Dr. E.R.C. Hagens, Department of Cranio- & Maxillofacial surgery, Visiting address: P. Debyelaan 25 | NL-6229 HX Maastricht Postal address: Postbus 5800, Tel: +31 (0)43 3872479; E-mail: eliza.hagens@mumc.nl
Abstract
Background: Tobacco, alcohol, and advanced age are well-established risk factors for oral cancer. However, a significant number of individuals develop oral cancer without these conventional risk factors. While the oral microbiome has been linked to cancers in other areas, its role as a risk factor for oral squamous cell carcinoma (OSCC) is unclear. This study aims to investigate the relationship between the oral microbiome and OSCC or premalignant conditions. Methods: proliferative verrucous hyperplasia, erosive lichen planus, submucous fibrosis) and 100 participants with healthy oral mucosa. Samples of buccal saliva and periodontal pockets will be collected. Samples of 25 healthy participants and 25 patients with OSCC will be collected at multiple time points (study day, 3-, 6- and 12 months after initial sampling or treatment for OSCC patients). Samples will be analysed using the interspacer profiling technique (ISpro) and Next Generation Sequencing (NGS). The primary outcome is the association between oral microbiome and (pre)malignant disease. Secondary outcomes include the association between oral microbiome and time, differences in microbiome be- tween different locations within patients and the changes of oral microbiome of healthy individuals over time and to assess whether the oral microbiome of individuals with OSCC changes after curative treatment. Discussion: Little is known about the association between OSCC and the oral microbiome. This investigation aims to elucidate this association, thereby enhancing our comprehension of OSCC etiology. Moreover, this study marks an initial stride towards developing a diagnostic tool for identifying individuals at heightened risk of OSCC. From a societal perspective, it addresses a significant health concern by potentially identifying the oral microbiome as an OSCC risk factor. This identification could facilitate early screening and preventive interventions, ultimately lowering cancer incidence and imparting a significant positive societal impact.
Trial registration
Keywords: oral microbiome, oral squamous cell carcinoma, oral cancer, head and neck cancer, cancer risk factors, oral flora
Background
Oral cavity cancer is a rare condition diagnosed approximately 3,000 times annually in the Netherlands and 380.000 worldwide. The majority of oral cavity cancers are squamous cell carcinomas (OSCC) [1,2]. While tobacco and alcohol use are recognized risk factors for OSCC, there is an observed increase in patients receiving this diagnosis despite an overall decline in the use of these substances [3]. Additionally, a subgroup of individuals presents with premalignant mucosal lesions. Significant resources are allocated to monitoring patients with premalignant lesions, yet only a small percentage (1-2%) progress to malignancy [4-6].
The mechanisms underlying the development of premalignant lesions and OSCC in patients with few or no identifiable risk factors remain unclear. Chronic inflammation is considered a potential contributing factor [7]. Periodontitis, affecting approximately 5-20% of the population, may play a role in the development of oral cavity cancer due to its persistent inflammatory status [8]. In vitro research has demonstrated that bacteria causing periodontitis, such as Porphyromonas gingivalis and Fusobacterium nucleatum, can stimulate certain cancer cells to undergo aggressive growth [9].Between 700-1,200 different species of bacteria cohabitate the human oral cavity, collectively forming together with numerous viruses and fungi the oral microbiome, unique to each individual [10,11]. Maintaining a harmonious balance between microorganisms, host cells, and the immune system ensures a symbiotic relationship. However, dysbiosis can disrupt this equilibrium, leading to various (systemic) diseases [12-14]. And has been linked to other types of including colorectal, pancreatic, and oesophageal cancers [15-17]. This high-lights the potential for microbiome-related mechanisms to play a broader oncogenic role, although the exact pathogenesis behind this process is not yet fully understood.
Few clinical studies have been conducted assessing differences in the oral micro-biome at the genus or phylum level between patients with and without head and neck squamous cell carcinoma (HNSCC). Although findings vary across studies, the most frequently reported observation is that samples from patients with OSCC and/or HNSCC are more often enriched in Prevotella and Fusobacterium [18-21]. However, results were conflicting. For instance, some studies report a lower abundance of streptococcus in OSCC patients, whereas others found higher levels in the same [20,21]. One study found that the commensal genera Corynebacterium and Kingella were associated with a lower risk of developing HNSCC. This study also suggests that their protective effect may be stronger in smokers, possibly due to the high-carcinogen environment in the oral cavity [22].
Two studies showed that even within the same patient, the oral microbiome differed significantly between tumour tissue and matched healthy sites. Tumour samples were enriched in periodontitis-associated taxa such as Fusobacterium, Dialister, Pepto-streptococcus, Filifactor, Peptococcus, Catonella, and Parvimonas [23,24].
Despite these findings, the current available literature are often inconsistent, how exactly the composition of oral microbiome differs from patients with OSCC and healthy individuals. With limited understanding of the underlying mechanisms by which the oral microbiome may influence carcinogenesis. A deeper understanding of these patterns is critical for advancing microbiome- informed cancer prevention or early detection strategies.
In addition to differences in prevalence between patients with and without OSCC, a Taiwanese study demonstrated that the abundance of specific bacteria increased progressively from stage I to stage IV [18,25]. Furthermore, microbiome differences have also been linked to other tumour characteristics, such as extranodal extension and the presence of lymph node metastases [26]. Besides differences between patients with OSCC and patients with a healthy oral mucosa, patients with premalignant lesions also have shown to have a significant difference in microbiome profiles. These microbiomes harbour less Streptococcus and show increase of fusobacterium and prevotella [20,26]. Moreover, little is known about the stability of the oral microbiome over time. A small longitudinal study showed that microbiome samples from saliva collected before and after surgery, alpha diversity decreased postoperatively but significantly increased in patients with cancer recurrence [18]. This suggests a potential relationship between microbiome diversity and cancer progression or relapse. The stability of the oral microbiome in healthy individuals remains unclear.
Important to note is that although the aforementioned studies provide valuable information, these studies are heterogeneous. Some focused solely on saliva samples while others took samples from (resected) specimens. Most studies were conducted in Asian populations, which differ from our Western populations in terms of genetic background, lifestyle, and environmental exposure, which is known to influence the oral microbiome [27]. Therefore, findings from these studies may not be generalizable to our local patient population, emphasizing the need for microbiome research in Western cohorts. Part of these studies had small sample sizes and did not assess the overall composition of the oral microbiome. Moreover, variables such as tumour stage (T stage), extranodal extension (ENE), oral health, smoking and alcohol consumption have been shown to significantly influence the oral microbiome, yet most of the studies did not take these variables into account in the data analyses [20,21,25,26].
It is hypothesized that the microbiome composition differs between patients with OSCC, patients with premalignant disease and healthy individuals. Furthermore, we hypothesize that the oral microbiome composition will show intraoral variability be-tween different sampling sites (e.g., buccal swap, periodontal pockets and tumour swab). Understanding these patterns could provide insights into the pathogenesis of OSCC and the potential diagnostic or prognostic utility of the oral microbiome.
The exact correlations between the composition of the oral microbiome and OSCC remains unclear. By analysing and comparing oral microbiome profiles for patients with OSCC, premalignant lesions and individuals with healthy mucosa, more will be known about the aetiology of OSCC and perhaps elucidate mechanisms underlying malignancies and premalignant conditions in patients, particularly those lacking typical risk factors. Ultimately this could lead to using (specific) changes in the oral micro- biome as a preventive screening tool and predictive indicator for the development of malignant abnormalities in the oral mucosa.
Taken together, these findings highlight the importance of conducting a large, longitudinal study in a representative patient population, using standardized methodologies and controlling for confounding factors. Such studies could contribute to clinical tools and interventions aimed at risk stratification and early cancer detection.
Therefore, the primary objective is to investigate whether the oral microbiome of individuals with oral squamous cell carcinoma (OSCC) differs from that of individuals with a healthy oral mucosa. As a control, we will longitudinally compare the oral microbiome among individuals with OSCC, to those with a premalignant lesion, and to those with a healthy oral mucosa. The secondary objective is to examine the characteristics of the oral microbiome. Therefore, we will assess differences in microbiome within individuals between different locations and whether and how the oral microbiome varies over time in individuals with a healthy oral mucosa and in individuals with OSCC who have undergone curative treatment.
Methods
Objective
The primary objective is to investigate whether the oral microbiome of individuals with oral squamous cell carcinoma (OSCC) differs from that of individuals with a healthy oral mucosa. As a control, we will compare the oral microbiome among individuals with OSCC, those with a premalignant lesion, and those with a healthy oral mucosa.
The secondary objective is to examine characteristics of the oral microbiome including differences in microbiome between different locations within patients, the as-sociation between the oral microbiome and age and the stability of the oral microbiome. Therefore, we will assess whether and how the oral microbiome varies over time in individuals with a healthy oral mucosa and in individuals with OSCC who have undergone curative treatment.
Study design and setting
In this prospective case-control study, patients visiting Maastricht University Medical Centre+ and Radboud University Medical Centre (The Netherlands) between December 2021 and January 2027 will be included.
Included patients are divided into three groups:
1. Patients without oral (pre)malignant conditions (n=100)
2. Patients with a premalignant* condition (n=100)
3. Patients with primary oral squamous cell carcinoma or verrucous carcinoma (n=100)
(* = erythroplakia, erythroleukoplakia, leukoplakia, proliferative verrucous hyperplasia, erosive lichen planus, submucosal fibrosis).
Multiple samples at different standardized location will be collected from each patient. Sample collection will be repeated for 50 patients after 3, 6 and 12 months. Afterwards, samples will be collected and analysed using the ISpro profiling technique and Next Generation Sequencing (NGS). The SPIRIT guidelines were used to write this protocol, ensuring transparent and complete reporting for this trial [28].
Study population
Patients are included if they meet the following criteria:
Inclusion
• Patients ≥18 years in case of oral malignancy or premalignant disease, ≥40 years in case of no malignant disease
• Patients are (partially) dentate (≥20 teeth)
• Patients who have given written informed consent to participate in this study
Exclusion
• Patients with a history of radiotherapy in head and neck region
• Patients with a history of chemotherapy, immunotherapy or immunomodulating drugs
• Patients with a previous malignancy in head and neck region
• Edentulous patients (with or without implants)
• Patients with removable (partial) dentures or occlusal splints
• Patients with solely lichen planus (and none of the aforementioned premalignant* diseases)
• Patients with braces
Inclusion and initial visit
Patients with OSCC are seen at the outpatient clinic following a standard head and neck care pathway. Patients are invited to participate in the study during one of the first visits in this head and neck care pathway. If they agree to participate, a pocket status is recorded during this initial appointment. During another standard appointment within the head and neck care pathway, samples are then collected.
Samples are taken at a subsequent visit, not simultaneously to avoid potential disruption of the microbiome during probing in pockets.
Healthy individuals and those with a premalignant condition are included from the outpatient clinic Cranio Maxillofacial Surgery, where they are asked to participate in the study during their first visit. Upon consent, a pocket measurement is conducted. Subsequently, samples are taken during the second (follow-up or treatment) appointment.
For the last 25 inclusions in the OSCC group and the group with healthy oral mucosa, patients are seen again after 3, 6, and 12 months to collect samples at those time points. Clinical data, such as patient characteristics, use of medication including antibiotics, tumour characteristics, type of premalignant condition, alcohol and smoking behaviour, and dental status, are also collected. The study procedures are depicted in Figure 1.
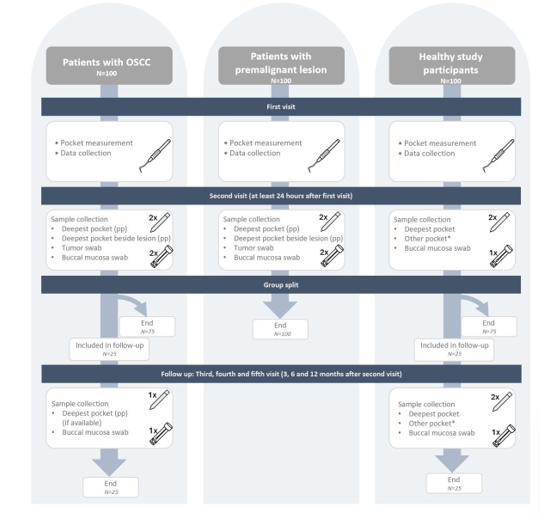
Figure 1:Flow chart study procedures
pp = paper point, * = Other pocket in another quadrant
Assessing gingival condition
In this study, we use the Periodontal Screening Score (PPS) to assess, standardize, and report the gingival condition of participants in a reproducible manner [31]. The PPS is a widely implemented and nationally recognized tool in the Netherlands for the early detection of periodontal disease. It provides a quick yet reliable screening of periodontal health and is routinely used in general dental practice.
The PPS was developed by the Dutch Association for Periodontology (NVvP) and is considered the national standard in periodontal screening [32]. It is based on a clinical examination using a periodontal probe and assigns a score per sextant ranging from score 1 (healthy gingiva) to score 4 (advanced periodontitis). Intermediate scores indicate increasing severity, and when a score of 3 or higher is recorded, a comprehensive periodontal assessment is typically recommended.
Sample collection procedure
All patients underwent a clinical examination at least one day prior to sample collection. During this initial visit a full periodontal assessment was conducted. During this examination, the depth and location of the deepest periodontal pocket in the oral cavity, the deepest pocket adjacent to the lesion, and a contralateral pocket were documented.
To allow the oral microbiome within the pockets sufficient time to stabilize following probing, sample collection was scheduled for a separate visit at least one day later. Multiple samples are taken from each patient at different locations to ensure the heterogeneity of microorganisms in the oral cavity and reduce sampling bias. Sterile paper points (PP) and sterile Floq swabs (swab) are used for this purpose. In all patients, a sample is taken from the deepest pocket (PP) and saliva from the buccal mucosa (swab). In individuals with healthy buccal mucosa, a third sample is obtained from a contralateral pocket (PP). In patients with malignancy or premalignant conditions, a third sample is obtained from the pocket next to the lesion (PP), and a fourth sample of saliva on the lesion (swab), table 1. In the 25 individuals with healthy buccal mucosa who are seen repeatedly for sample collection, only the deepest pocket and buccal mucosa (swab) will be sampled during the measurements after 3, 6 and 12 months. In the 25 patients with OSCC, the repeated samples will be collected from the deepest pocket (if still available after surgery) and the buccal swab. Sample collection in done by one of the oral hygienist or researchers using standardised instructions, appendix 1.
Table 1. Sample locations for different participant groups
| Patients with OSCC | Patients with premalignant disease | Individuals with healthy oral mucosa |
|---|---|---|
| (1) Deepest pocket (PP) As determined during previous visit | (1) Deepest pocket (PP) As determined during previous visit | (1) Deepest pocket (PP) As determined during previous visit |
| (2) Deepest pocket near tumor (PP) As determined during previous visit | (2) Deepest pocket near tumor (PP) As determined during previous visit | (2) Other pocket (PP)* As determined during previous visit |
| (3) Tumor (swab) | (3) Tumor (swab) | (3) Healthy buccal mucosa (swab) |
| (4) Healthy buccal mucosa (swab) | (4) Healthy buccal mucosa (swab) |
* = random other pocket other quadrant, pp = paper point, swab = floq swab.
Sample analysis
The gold standard for microbial identification is laboratory cultivation. However, a portion of oral cavity microorganisms cannot be cultured. To identify all bacteria in the oral microbiome, the Polymerase Chain Reaction (PCR) profiling technique IS-pro is employed. IS-pro enables the identification of bacteria by amplifying the 16S-23S interspace region in bacterial DNA. The length of each amplified IS region is species specific and is used to identify the bacterial species present in the sample [29,30]. This procedure is conducted by trained research technicians. Research technicians were blinded to both the sample location and the study group.
Treatment of patients with OSCC
Usually, patients with a suspicious lesion of the oral mucosa are staged with biopsies of the primary tumour followed by imaging, typically using Magnetic Resonance Imaging (MRI) and/or computed tomography (CT), to assess the extent of invasion into surrounding tissues. Fine Needle Aspiration Cytology (FNAC) is used for cN staging. A conventional chest X-ray is used for cM staging unless there are medical reasons for a chest CT. Additionally, all patients with OSCC are referred to the CBT (Center for Special Dental Care, Centrum Bijzondere Tandheelkunde) for a thorough dental ex-amination, including an Orthopantomogram (OPT), to identify any potential dental foci that may complicate treatment.
The primary curative treatment consists of surgical resection with or without adjuvant radiotherapy and/or chemotherapy. Due to a close anatomical relation of the oral mucosa to the bone of the maxilla and mandibula bone invasion often occurs. The amount of invasion guides the bony resection margins. A marginal mandibular resection is performed for patients with limited cortical bone involvement, where an adequate surgical bone margin can be achieved. For patients with more extensive bone invasion, where an adequate oncologic bone margin cannot be obtained or insufficient mandibular bone remains to ensure stability, a segmental resection is indicated. For patients with a small OSCC (cT1-T2) and a clinically negative neck (cN0), options include elective neck dissection, a sentinel lymph node biopsy, or a watch-and-wait strategy with regular ultrasonographic monitoring of the neck. Each approach aims to manage the potential spread of cancer while minimizing unnecessary surgical intervention. Patients with >T2 and/or N+ staging undergo a elective neck dissection on the ipsilateral and/or contralateral side depending on the tumour location. Defects are re-constructed primarily, with local flaps or free flap transferred tissue. All patients are discussed in a head and neck multidisciplinary team (MDT) pre- and postoperatively and treated according the Dutch National Guidelines in Head and Neck Cancer.33 The risk of recurrence is estimated combining surgical margins and the presence of lymphatic and/or vascular invasion and growth pattern: high risk is defined as less than 1mm, intermediate risk is 1 to 4mm, and low risk is 5 mm or more. Low risk patients only receive clinical follow up, intermediate risk margins may require adjuvant treatment, either through radiotherapy or surgical re-resection. High-risk patients are presented for radiotherapy and/or chemotherapy depending on their co-morbidities and age. All patients are in a follow up protocol for at least 5 years post-treatment.
Primary endpoint
The primary outcome is the composition of the oral microbiome for patients with OSCC, patients with premalignant disease and individuals with healthy oral mucosa. Microbiome composition will be compared between these groups on both phyla level and species level. These analyses will be performed for both samples from pockets and buccal saliva separately.
Secondary endpoints
• Examine characteristics of the oral microbiome
o Differences between microbiome composition within individuals between different sample locations.
o Assess if the oral microbiome varies over time in individuals with a healthy oral mucosa.
o Assess if the oral microbiome varies in patients with OSCC after they have undergone curative treatment.
o Assess if there are significant association between oral microbiome and age.
Statistical analysis
Baseline characteristics such as age, gender, comorbidities, tobacco- and alcohol use, toothbrushing frequency, use of antibiotics or anti-inflammatory medication in the last month, use of chlorhexidine mouthwash in the last week, presence of deep caries or retained root(s), depth of periodontal pockets and tumour characteristics will be dis-played to give insight in the type of patients included in this study. Data will be checked for normality graphically by using a histogram and Q - Q plot. Continuous variables with normal distributions will be presented as means with standard deviations and will be compared using independent t-test. Medians and interquartile ranges (IQR) will be used as central tendency for continuous variables with non-normal distributions, these data will be compared using the Mann–Whitney U-test. Categorical data will be expressed with percentage frequencies and will be compared using a Chi-square or Fisher’s exact test where appropriate. To investigate the structure of the oral microbiome, we will evaluate α-diversity (which reflects the diversity within a single sample) and β-diversity which measures differences in microbial composition between samples, e.g., between individuals and between sampling locations). β-diversity will be assessed using multiple distance metrics, including Bray-Curtis and cosine similarity. To statistically evaluate group-level differences in β-diversity, we will use Permutational Multivariate Analysis of Variance (PERMANOVA).
Differential abundance of microbial taxa between groups (e.g., OSCC, premalignant lesions, and healthy mucosa) will be assessed at both the phylum and species levels. Analyses will be conducted both within individuals (e.g., between sample sites or across time points) and between groups. To account for multiple comparisons across taxa, p-values will be corrected using the Benjamini-Hochberg false discovery rate (FDR) correction.
Potential confounders such as age, sex, tobacco use, alcohol use, comorbidities, T-stage, PPS score and oral hygiene practices will be included as covariates in multi-variable models to adjust for their influence on microbiome composition.
To analyse longitudinal changes in the oral microbiome within individuals, linear mixed-effects models will be used with time point as a fixed effect and individual as a random effect. Potential confounders will be incorporated in the statistical models as fixed effects.
No formal power analysis was performed, as no comparable prior studies or pilot data were available to inform accurate effect size estimates. This exploratory study aims to provide the necessary data to enable future sample size calculations.
Discussion
In this case control study, we aim to assess the association between the oral microbiome and OSCC. To date, little is known about the oral microbiome and how to study it. For instance, it is unknown what is the best way or location is to obtain samples. There is no answer to how much does the oral microbiome vary between sample location within individuals. Little is known about the stability of the oral microbiome. And finally, it remains unclear if the oral microbiome changes over time. These are factors that are also relevant to the primary aim of this study, as these factors influence the possible predictive value of the oral microbiome for OSCC. To address this gap in knowledge, secondary outcomes have been established to assess these uncertainties.
This study presents an innovative approach to investigating the relationship between the oral microbiome and OSSC. This study includes not only patients with OSCC, but also individuals with histologically confirmed premalignant lesions, allowing for a broader understanding of microbial changes. Furthermore, the sampling strategy is feasible and translatable to the clinical setting. Also, the microbiome samples are collected from multiple intraoral sites, including tumor swab, contralateral healthy mucosa swab, and samples from subgingival pockets. This enables us to assess the difference between sample locations and potentially identify the optimal sample location. The inclusion of pocket samples is particularly noteworthy, as it provides insight into the potential role of periodontal health and microbiome in OSCC.
Additionally, this study evaluates the composition of the oral microbiome over time in both OSCC patients and healthy individuals. Little previous research has been done on this topic previously, while this is relevant for future diagnostic or prognostic applications. In the current study factors such as antibiotic use, smoking, alcohol consumption, oral health status, and tumor characteristics are also taking into account. This strengthens the reliability and clinical relevance of the study findings.
Together, these elements make this study an innovation contribution to the field, with potential implications for both scientific understanding and clinical translation in the context of oral cancer. This study can be the basis for follow-up studies aiming to develop tools for risk stratification strategies or early detection of OSCC. Furthermore, if distinct microbial patterns are identified in this study, follow- up research may help to clarify potential causal. For these studies functional omics approaches, such as meta-genomics and proteomics, could be used to investigate the metabolic activity and mechanistic contribution of bacteria, and also fungi and viruses in oral carcinogenesis.
The present study will provide crucial insights into the role of the oral microbiome in OSCC development. It has the potential to create a reliable diagnostic tool for timely identification of high- risk individuals, improving early detection and treatment. By involving primary care providers, it aims to reduce the growing cases of oral cavity cancer, offering less invasive treatments and enhancing the overall quality of life for patients.
Trial Status
The current approved protocol version is 2.0 (version date 16-08-2023). Patient enrollment began 1st December 2021 and we anticipate recruitment to be finished on 1st January 2027.
Abbreviations
OSCC: Oral squamous cell carcinoma; PCR: Polymerase Chain Reaction; NGS: Next Generation Sequencing; PP: Paperpoint; MRI: Magnetic Resonance Imaging; CT: Computed Tomography; OPT: Orthopantomogram; FNAC: Fine Needle Aspiration Cytology; CBT: Centrum Bijzondere Tandheelkunde (Center for Special Dental Care); IQR: Interquartile range; PLS-DA: Partial least squares discriminant analysis
Declarations
Ethics approval and consent to participate
Approval of Medical Research Ethics Committees of both Maastricht UMC and Radboud UMC had been obtained July 9th 2021 and amendment to current protocol on August 16th 2023 under approval number METC-2019-1251-A-11.
Consent for publication
Not applicable.
Availability of data and materials
The final trial dataset will be accessible for solely the investigators.
Competing interests
The authors declare that they have no competing interests.
Funding
Funding has been obtained from Wetenschappelijke Onderwijs- en Onderzoeksactiviteiten (BOOA), Academische Alliantie Fonds (Radboudumc en Maastricht UMC+) and crowdfunding. The funding bodies did not have any role in the design of the study and will not have any role in collection, analysis, and interpretation of data and in writing the manuscript.
Authors' contributions
Conceptualization, L.V. and .; methodology, L.V., E.H., P.K., P.S..; writing—original draft preparation, E.H., L.V..; writing—review and editing, E.H.,.; visualization, E.H., L.V. and C.K..; supervision, L.V., P.K., P.S..; project administration, C,K, and E.H..; funding acquisition, L.V., E.H. and C.K.. All authors have read and agreed to the published version of the manuscript.
Acknowledgements
Not applicable.
References
1. Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209-49.
2. NKR cijfers. IKNL. Available from: https://iknl.nl/nkr-cijfers. Accessed 2023 Mar 10.
3. Tran Q, Maddineni S, Arnaud EH, Divi V, Megwalu UC, Topf MC, et al. Oral cavity cancer in young, non-smoking, and non-drinking patients: a contemporary review. Crit Rev Oncol Hematol. 2023;190:104112.
4. Guan JY, Luo YH, Lin YY, Wu ZY, Ye JY, Xie SM, et al. Malignant transformation rate of oral leukoplakia in the past 20 years: a systematic review and meta-analysis. J Oral Pathol Med. 2023;52(1):1-10.
5. Evren I, Brouns ER, Wils LJ, Poell JB, Peeters CFW, Brakenhoff RH, et al. Annual malignant transformation rate of oral leukoplakia remains consistent: a long-term follow-up study. Oral Oncol. 2020;110:104860.
6. Muller S, Tilakaratne WM. Update from the 5th edition of the World Health Organization classification of head and neck tumors: tumours of the oral cavity and mobile tongue. Head Neck Pathol. 2022;16(1):54-62.
7. Radaic A, Ganther S, Kamarajan P, Grandis J, Yom SS, Kapila YL. Paradigm shift in the pathogenesis and treatment of oral cancer focused on the oralome and antimicrobial-based therapeutics. Periodontol 2000. 2021;87(1):76-93.
8. Frencken JE, Sharma P, Stenhouse L, Green D, Laverty D, Dietrich T. Global epidemiology of dental caries and severe periodontitis: a comprehensive review. J Clin Periodontol. 2017;44(S18):S94-105.
9. Nagy KN, Sonkodi I, Szöke I, Nagy E, Newman HN. The microflora associated with human oral carcinomas. Oral Oncol. 1998;34(4):304-8.
10. Wade WG. The oral microbiome in health and disease. Pharmacol Res. 2013;69(1):137-43.
11. Dewhirst FE, Chen T, Izard J, Paster BJ, Tanner ACR, Yu WH, et al. The human oral microbiome. J Bacteriol. 2010;192(19):5002-17.
12. Sanz M, Del Castillo AM, Jepsen S, Gonzalez-Juanatey JR, D'Aiuto F, Bouchard P, et al. Periodontitis and cardiovascular diseases: consensus report. Glob Heart. 2020;15(1):1-34. doi:10.5334/gh.400.
13. Sanz M, Ceriello A, Buysschaert M, Chapple I, Demmer RT, Graziani F, et al. Scientific evidence on the links between periodontal diseases and diabetes: consensus report of the joint workshop. J Clin Periodontol. 2018;45(2):138-49.
14. Herrera D, Sanz M, Shapira L, Brotons C, Chapple I, Frese T, et al. Association between periodontal diseases and cardiovascular, diabetes and respiratory diseases: consensus report. J Clin Periodontol. 2023;50(7):819-41.
15. Li X, Zhu S, Zhang T, Chen X. Association between oral microflora and gastrointestinal tumors. Oncol Rep. 2021;46(2):160.
16. Farrell JJ, Zhang L, Zhou H, Chia D, Elashoff D, Akin D, et al. Variations of oral microbiota are associated with pancreatic diseases including pancreatic cancer. Gut. 2012;61(4):582-8.
17. Zepeda-Rivera M, Minot SS, Bouzek H, Wu H, Blanco-Míguez A, Manghi P, et al. A distinct Fusobacterium nucleatum clade dominates the colorectal cancer niche. Nature. 2024;628(8007):424-32.
18. Guerrero-Preston R, Godoy-Vitorino F, Jedlicka A, Rodríguez-Hilario A, González H, Bondy J, et al. 16S rRNA amplicon sequencing identifies microbiota associated with oral cancer and HPV. Oncotarget. 2016;7(32):51320-34.
19. Zhang L, Liu Y, Zheng HJ, Zhang CP. The oral microbiota may have influence on oral cancer. Front Cell Infect Microbiol. 2020;9:476.
20. Yang CC, Washio J, Lin YC, Hsu ML, Wang DH, Tsai FT, et al. Microbiome signatures and dysbiotic patterns in oral cancer. Oral Dis. 2025 Mar 19.
21. Chan JYK, Cheung MK, Lan L, Ng C, Lau EHL, Yeung ZWC, et al. Characterization of oral microbiota in HPV and non-HPV head and neck SCC. Oral Oncol. 2022;135:106245.
22. Hayes RB, Ahn J, Fan X, Peters BA, Ma Y, Yang L, et al. Association of oral microbiome with risk for incident HNSCC. JAMA Oncol. 2018;4(3):358-65.
23. Zhao H, Chu M, Huang Z, Yang X, Ran S, Hu B, et al. Variations in oral microbiota associated with oral cancer. Sci Rep. 2017;7:11773.
24. Schmidt BL, Kuczynski J, Bhattacharya A, Huey B, Corby PM, Queiroz EL, et al. Changes in abundance of oral microbiota associated with oral cancer. PLoS One. 2014;9(6):e98741.
25. Yang CY, Yeh YM, Yu HY, Chin CY, Hsu CW, Liu H, et al. Oral microbiota community dynamics associated with OSCC staging. Front Microbiol. 2018;9:862.
26. Zeng B, Tan J, Guo G, Li Z, Yang L, Lao X, et al. The oral cancer microbiome contains tumor space-specific bacteria. Front Cell Infect Microbiol. 2022;12:942328.
27. Stankevic E, Kern T, Borisevich D, Poulsen CS, Madsen AL, Hansen TH, et al. GWAS identifies host genetic variants influencing oral microbiota diversity. Sci Rep. 2024;14(1):14738.
28. Chan AW, Tetzlaff JM, Gøtzsche PC, Altman DG, Mann H, Berlin JA, et al. SPIRIT 2013 explanation and elaboration. BMJ. 2013;346:e7586.
29. Budding AE, Grasman ME, Lin F, Bogaards JA, Soeltan-Kaersenhout DJ, Vandenbroucke-Grauls CM, et al. IS-pro: high-throughput molecular fingerprinting of the intestinal microbiota. FASEB J. 2010;24(11):4556-64.
30. Inbiome. IS-pro profiling technique. Available from: http://isdiagnostics.nl/technology. Accessed 2023 Mar 10.
31. Louropoulou A, van der Weijden F. De richtlijnen voor parodontale en peri-implantaire ontstekingen. Ned Tijdschr Tandheelkd. 2021;128(12):585-92. Dutch.
32. Nederlandse Vereniging voor Parodontologie. Available from: https://www.nvvp.org/home. Accessed 2025 Jul 18.
33. Federatie Medisch Specialisten. Hoofd-halstumoren. Richtlijnendatabase. Available from: https://richtlijnendatabase.nl/richtlijn/hoofd-halstumoren. Accessed 2024 Feb 23.
Received: October 31, 2025;
Accepted: November 17, 2025;
Published: November 26, 2025.
To cite this article : Hagens ERC, Kiewiet CC, Kessler PAWH, Coppen C, Savelkoul PHM, Vaassen LAA. Comparison of oral microbiome in patients with and without oral squamous cell carcinoma: study protocol for a prospective case-control trial. Br J Cancer Res. 2025;8(3):832-839. doi:10.31488/bjcr.212.
© The Author(s) 2025. This is an open access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/).
